Empirical Methods in Finance
Part 1
Agenda
Agenda
- Apresentação do syllabus do curso
- Apresentação dos critérios de avaliação
- Breve apresentação dos temas de pesquisa e discussão inicial sobre a entrega final
- Início do conteúdo
- Introdução a causalidade
Stata
Providenciar instalação para próximo encontro.
Para instalação do Stata, seguir instruções da TI.
R
Providenciar instalação para próximo encontro.
Install R here Win
Install R here Mac
Install R Studio here
Para instalar e carregar os pacotes você precisa rodar as duas linhas abaixo.
Python
I might show some code in python, but I cannot offer you support on it.
Selection bias

Você nunca sabe o resultado do caminho que não toma.
Quais as aplicações do que vamos discutir?
Há uma série de questões de pesquisa que poderiam ser investigadas com as ferramentas que vamos discutir hoje.
Vale mais a pena estudar em escola particular ou pública?
Qual o efeito de investimentos de marketing têm na lucratividade?
Qual o efeito que jornadas de 4 dias semanais têm na produtividade?
Qual efeito que educação tem na remuneração futura?
E diversas outras semelhantes…
Antes de começar: Nossa agenda
Introdução a pesquisa quantitativa
Validade Externa vs. Validade Interna
Problemas em pesquisa quantitativa inferencial
Remédios
Introdução
O que fazemos em pesquisa quantitiva? Seguimos o método de pesquisa tradicional (com ajustes):
Observação
Questão de pesquisa
Modelo teórico (abstrato)
Hipóteses
Modelo empírico
Coleta de dados
Análise do resultado do modelo (diferente de análise de dados “pura”)
Conclusão/desdobramentos/aprendizados
Introdução
O que fazemos em pesquisa quantitiva? Seguimos o método de pesquisa tradicional (com ajustes):
Observação
Questão de pesquisa
Modelo teórico (abstrato): Aqui é onde a matemática é necessária
Hipóteses
Modelo empírico: Estatística e econometria necessárias
Coleta de dados: Geralmente secundários
Análise do resultado do modelo (diferente de análise de dados “pura”)
Conclusão/desdobramentos/aprendizados
Definição
Pesquisa quantitativa busca testar hipóteses…
…a partir da definição de modelos formais (abstratos)…
…de onde se estimam modelos empíricos utilizando a estatística e a econometria como mecanismos/instrumentos.
No fim do dia, buscamos entender as relações (que tenham validade interna e que ofereçam validade externa) entre diferentes variáveis de interesse.
Quais as vantagens?
- Validade externa:
- Conceito de que, se a pesquisa tem validade externa, os seus achados são representativos.
- I.e., são válidos além do seu modelo. Resultados “valem externamente”.
- Idealmente, buscamos resultados que valem externamente para acumular conhecimento…
- …naturalmente, nem toda pesquisa quantitativa oferece validade externa. A pesquisa ótima sim. A pesquisa excelente tem validade externa para além do seu tempo.
- Pesquisa qualitativa dificilmente oferece validade externa.
Quais as armadilhas?
- Validade interna:
- Conceito de que a pesquisa precisa de validade interna para que seus resultados sejam críveis.
- I.e., os resultados não podem conter erros, vieses, problemas de estimação, problemas nos dados, etc..
- É aqui que a gente separa a pesquisa ruim da pesquisa boa. Para ser levada a sério, a pesquisa PRECISA ter validade interna.
- Mas isso, nem sempre é trivial. Muitas pesquisas que vemos publicadas, mesmo em top journals, não têm validade interna (seja por erro do pesquisador, por método incorreto, por falta de dados…)
- Mas cada vez mais, avaliadores estão de olho em problemas e em modelos Trash-in-Trash-out
Como fazemos na prática?
Exemplo de modelo empírico:
\(Y_{i} = α + 𝜷_{1} × X_i + Controls + error\)
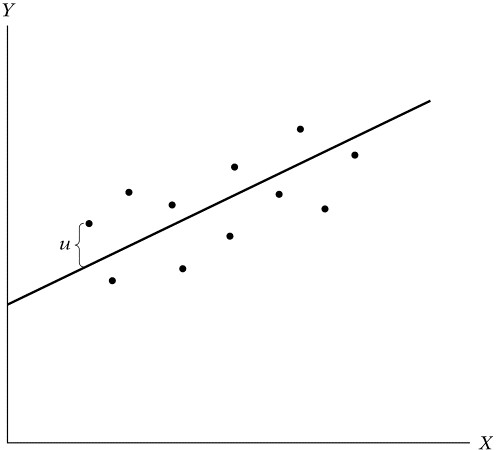
Uma vez que estimemos esse modelo, temos o valor, o sinal e a significância do \(𝜷\).
Se o Beta for significativamente diferente de zero e positivo –> X e Y estão positivamente correlacionados.
O problema? Os pacotes estatísticos que utilizamos sempre “cospem” um beta. Seja ele com ou sem viés.
Cabe ao pesquisador ter um design empírico que garanta que o beta estimado tenha validade interna.
Como fazemos na prática?
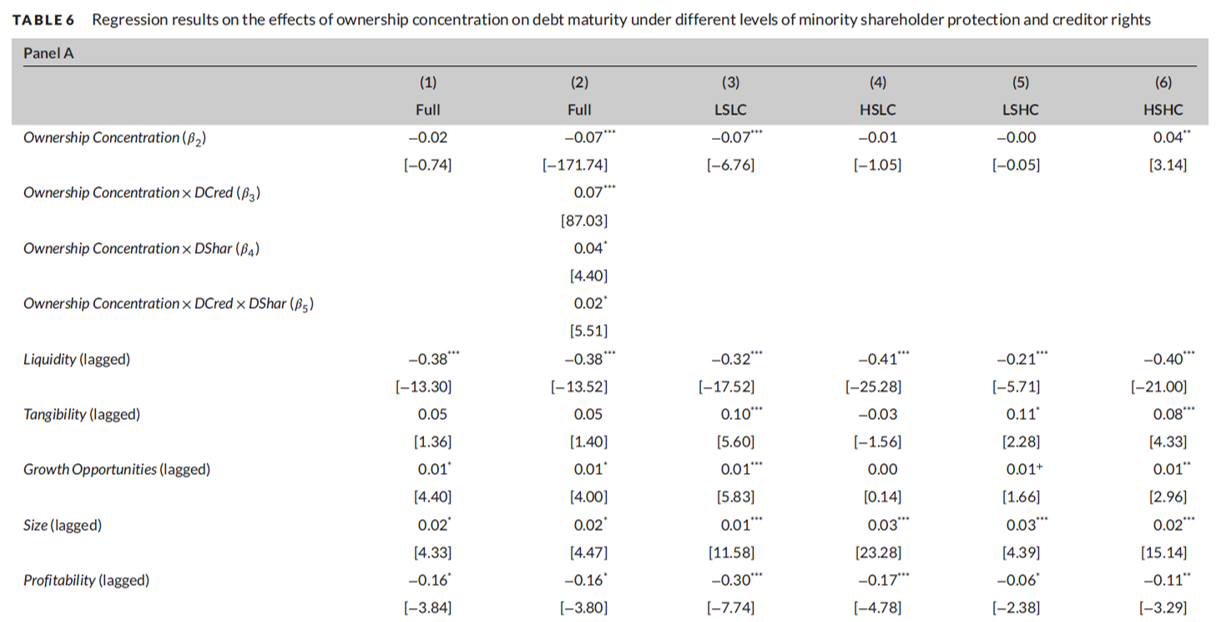
A decisão final é baseada na significância do Beta estimado. Se significativo, as variáveis são relacionadas e fazemos inferências em cima disso.
Contudo, sem um design empírico inteligente, o beta encontrado pode ter literalmente qualquer sinal e significância.
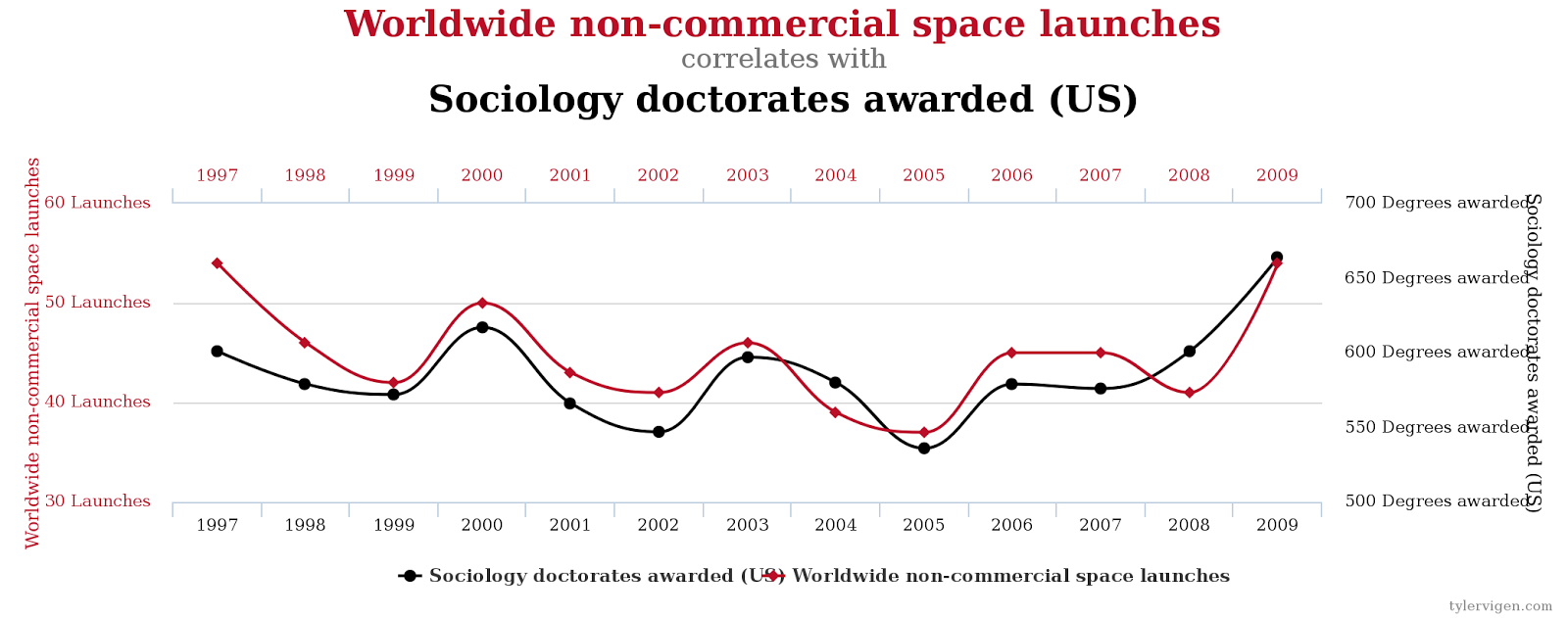
Exemplo desses problemas
Veja esse site.
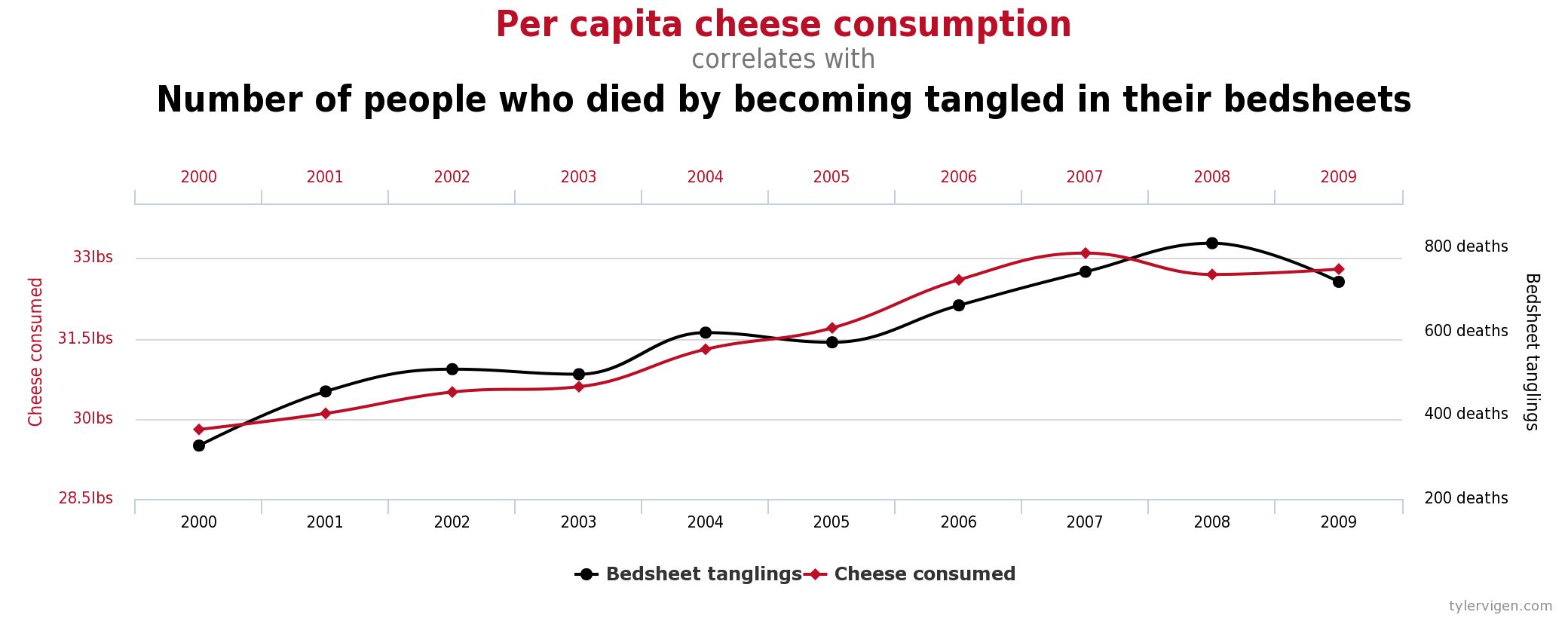
Exemplo desses problemas
Veja esse site.
Selection bias - We see I
R
library(data.table)
library(ggplot2)
# Generate Data
n = 10000
set.seed(100)
x <- rnorm(n)
y <- rnorm(n)
data1 <- 1/(1+exp( 2 - x - y))
group <- rbinom(n, 1, data1)
# Data Together
data_we_see <- subset(data.table(x, y, group), group==1)
data_all <- data.table(x, y, group)
# Graphs
ggplot(data_we_see, aes(x = x, y = y)) +
geom_point(aes(colour = factor(-group)), size = 1) +
geom_smooth(method=lm, se=FALSE, fullrange=FALSE)+
labs( y = "", x="", title = "The observations we see")+
xlim(-3,4)+ ylim(-3,4)+
theme(plot.title = element_text(color="black", size=30, face="bold"),
panel.background = element_rect(fill = "grey95", colour = "grey95"),
axis.text.y = element_text(face="bold", color="black", size = 18),
axis.text.x = element_text(face="bold", color="black", size = 18),
legend.position = "none")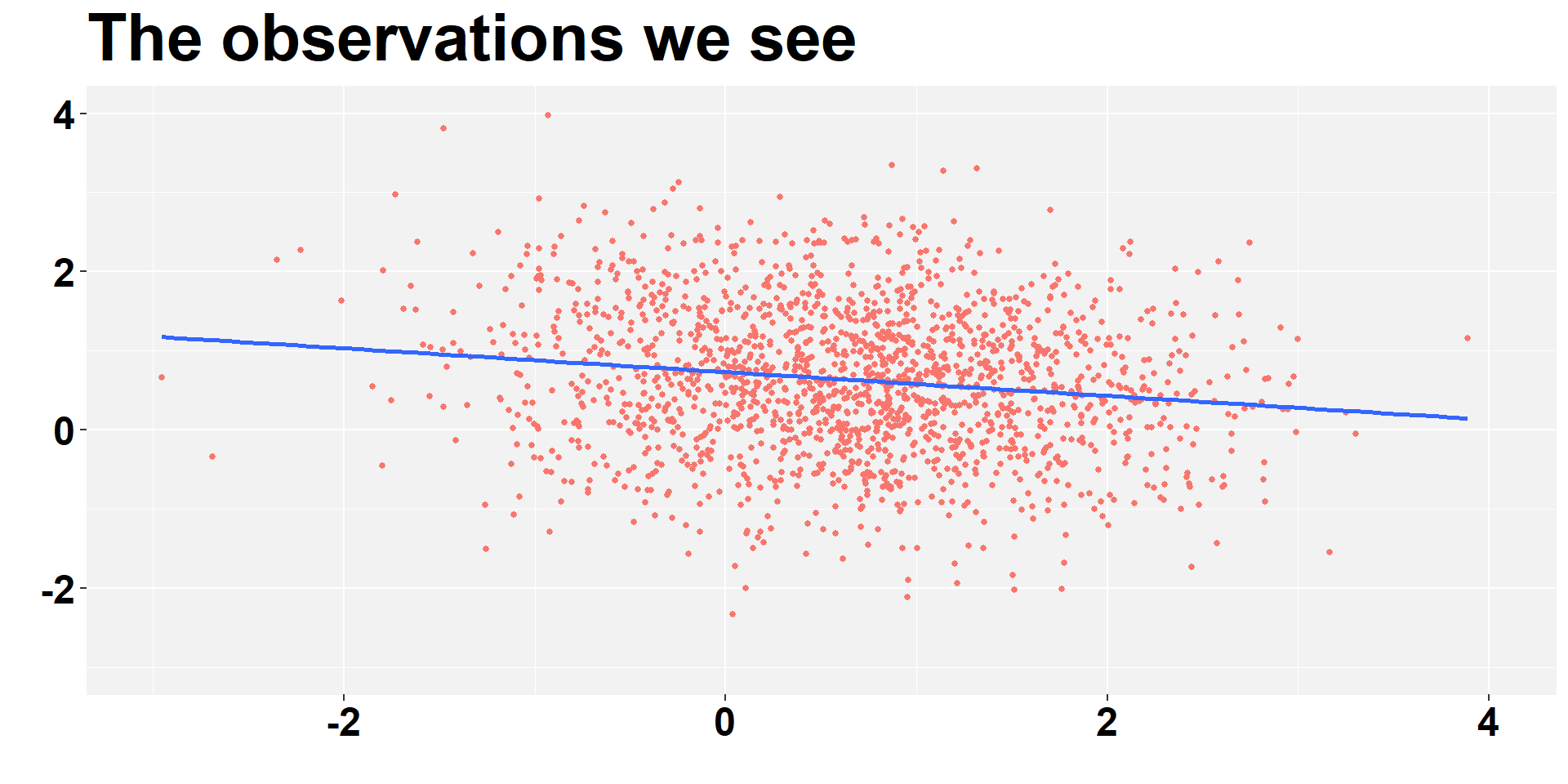
Python
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
n = 10000
np.random.seed(100)
x = np.random.normal(size=n)
y = np.random.normal(size=n)
data1 = 1 / (1 + np.exp(2 - x - y))
group = np.random.binomial(1, data1, n)
data_we_see = pd.DataFrame({'x': x[group == 1], 'y': y[group == 1], 'group': group[group == 1]})
data_all = pd.DataFrame({'x': x, 'y': y, 'group': group})
sns.set(style='whitegrid')
plt.figure(figsize=(7, 5))
plt.scatter(data_we_see['x'], data_we_see['y'], c=-data_we_see['group'], cmap='viridis', s=20)
sns.regplot(x='x', y='y', data=data_we_see, scatter=False, ci=None, line_kws={'color': 'blue'})
plt.title("The observations we see", fontsize=18)
plt.xlabel("")
plt.ylabel("")
plt.show()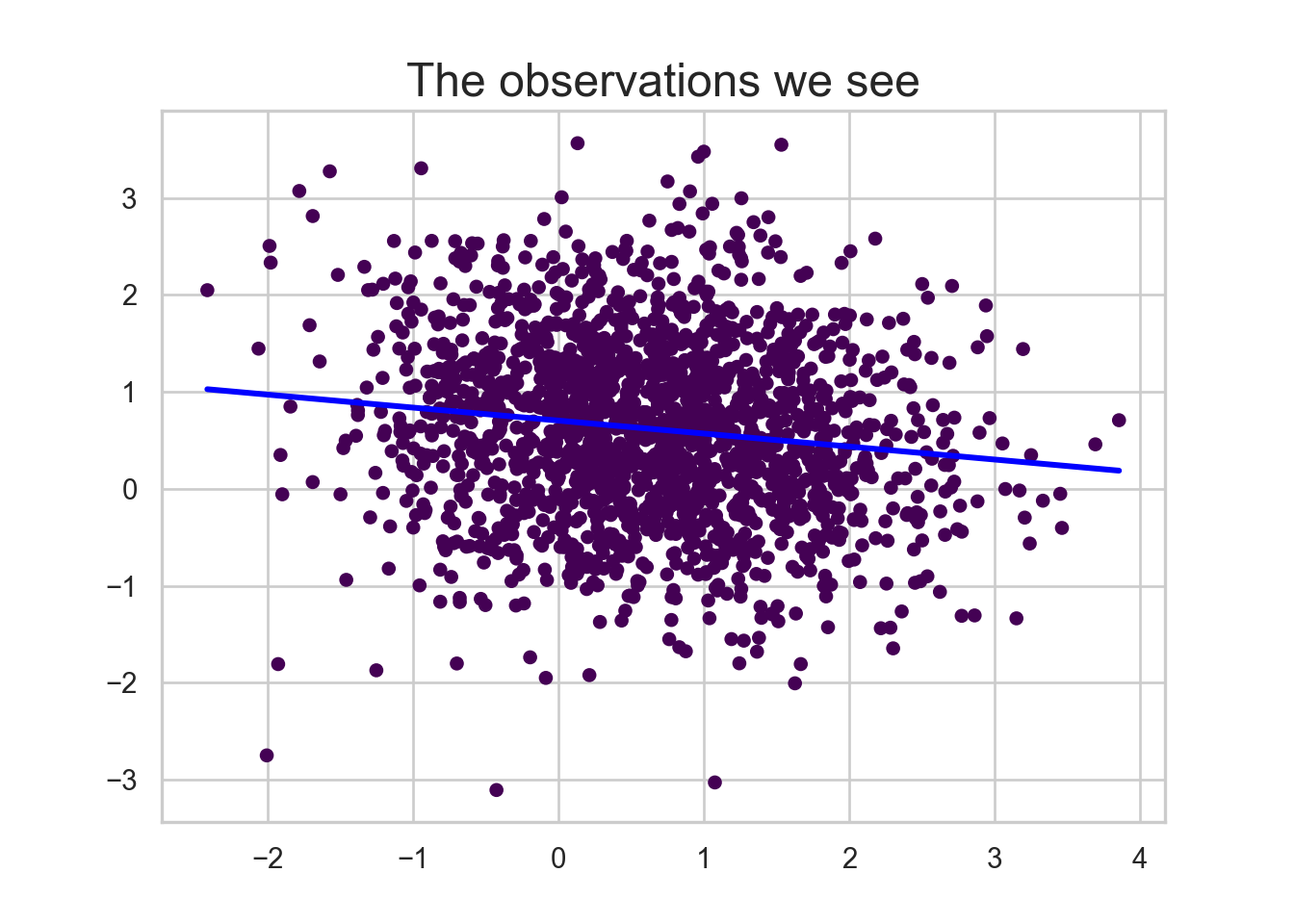
Stata
clear all
set seed 100
set obs 10000
gen x = rnormal(0,1)
gen y = rnormal(0,1)
gen data1 = 1 / (1 + exp(2 - x - y))
gen group = rbinomial(1, data1)
twoway (scatter x y if group == 1, mcolor(black) msize(small)) (lfit y x if group == 1, color(blue)),title("The observations we see", size(large) ) xtitle("") ytitle("")
quietly graph export figs/graph1.svg, replaceSelection bias - We see II
R
Call:
lm(formula = y ~ x, data = data_we_see)
Residuals:
Min 1Q Median 3Q Max
-3.05878 -0.63754 -0.00276 0.62056 3.11374
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.72820 0.02660 27.37 < 2e-16 ***
x -0.14773 0.02327 -6.35 2.75e-10 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.9113 on 1747 degrees of freedom
Multiple R-squared: 0.02256, Adjusted R-squared: 0.022
F-statistic: 40.32 on 1 and 1747 DF, p-value: 2.746e-10Python
import statsmodels.api as sm
import pandas as pd
n = 10000
np.random.seed(100)
x = np.random.normal(size=n)
y = np.random.normal(size=n)
data1 = 1 / (1 + np.exp(2 - x - y))
group = np.random.binomial(1, data1, n)
data_we_see = pd.DataFrame({'x': x[group == 1], 'y': y[group == 1], 'group': group[group == 1]})
data_all = pd.DataFrame({'x': x, 'y': y, 'group': group})
X = data_we_see['x']
X = sm.add_constant(X)
y = data_we_see['y']
model = sm.OLS(y, X).fit()
print(model.summary()) OLS Regression Results
==============================================================================
Dep. Variable: y R-squared: 0.018
Model: OLS Adj. R-squared: 0.018
Method: Least Squares F-statistic: 33.84
Date: sex, 29 ago 2025 Prob (F-statistic): 7.06e-09
Time: 13:31:25 Log-Likelihood: -2411.1
No. Observations: 1809 AIC: 4826.
Df Residuals: 1807 BIC: 4837.
Df Model: 1
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
const 0.7037 0.026 26.826 0.000 0.652 0.755
x -0.1339 0.023 -5.817 0.000 -0.179 -0.089
==============================================================================
Omnibus: 4.656 Durbin-Watson: 1.973
Prob(Omnibus): 0.097 Jarque-Bera (JB): 5.264
Skew: -0.038 Prob(JB): 0.0720
Kurtosis: 3.253 Cond. No. 1.93
==============================================================================
Notes:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.Stata
Number of observations (_N) was 0, now 10,000.
Source | SS df MS Number of obs = 1,872
-------------+---------------------------------- F(1, 1870) = 48.62
Model | 40.9398907 1 40.9398907 Prob > F = 0.0000
Residual | 1574.57172 1,870 .842016963 R-squared = 0.0253
-------------+---------------------------------- Adj R-squared = 0.0248
Total | 1615.51161 1,871 .863448215 Root MSE = .91761
------------------------------------------------------------------------------
y | Coefficient Std. err. t P>|t| [95% conf. interval]
-------------+----------------------------------------------------------------
x | -.1579538 .0226526 -6.97 0.000 -.2023808 -.1135269
_cons | .7202285 .0257215 28.00 0.000 .6697827 .7706744
------------------------------------------------------------------------------Selection bias - All I
R
ggplot(data_all, aes(x = x, y = y, colour=group)) +
geom_point(aes(colour = factor(-group)), size = 1) +
geom_smooth(method=lm, se=FALSE, fullrange=FALSE)+
labs( y = "", x="", title = "All observations")+
xlim(-3,4)+ ylim(-3,4)+
theme(plot.title = element_text(color="black", size=30, face="bold"),
panel.background = element_rect(fill = "grey95", colour = "grey95"),
axis.text.y = element_text(face="bold", color="black", size = 18),
axis.text.x = element_text(face="bold", color="black", size = 18),
legend.position = "none")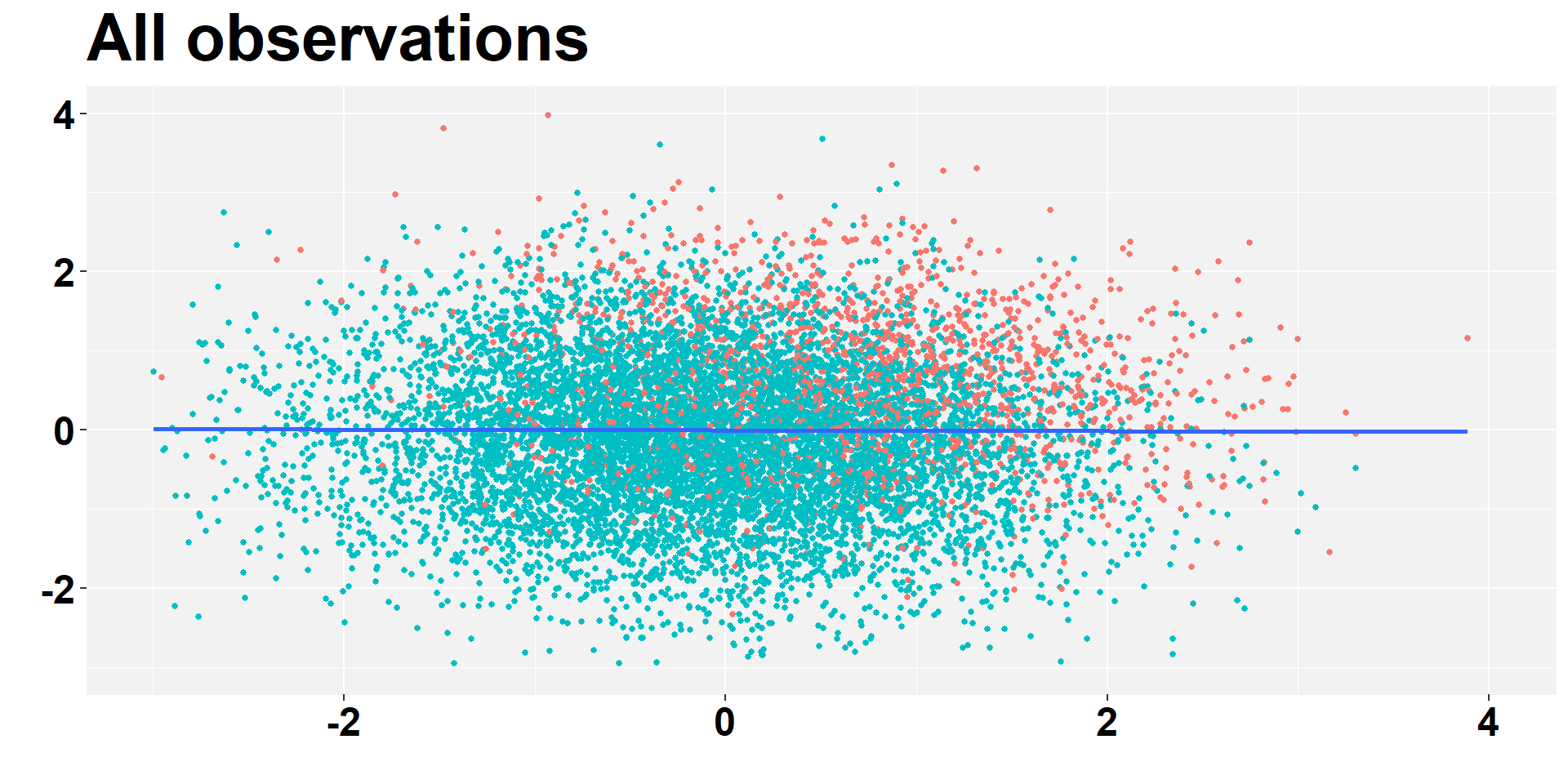
Python
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style='whitegrid')
plt.figure(figsize=(6, 4))
sns.scatterplot(data=data_all, x='x', y='y', hue='group', palette=['blue', 'red'], s=20)
sns.regplot(data=data_all, x='x', y='y', scatter=False, ci=None, line_kws={'color': 'blue'})
plt.title("All observations", fontsize=18)
plt.xlabel("")
plt.ylabel("")
plt.legend(title="Group", labels=["0", "1"], loc="upper left")
plt.gca().get_legend().remove()
plt.show()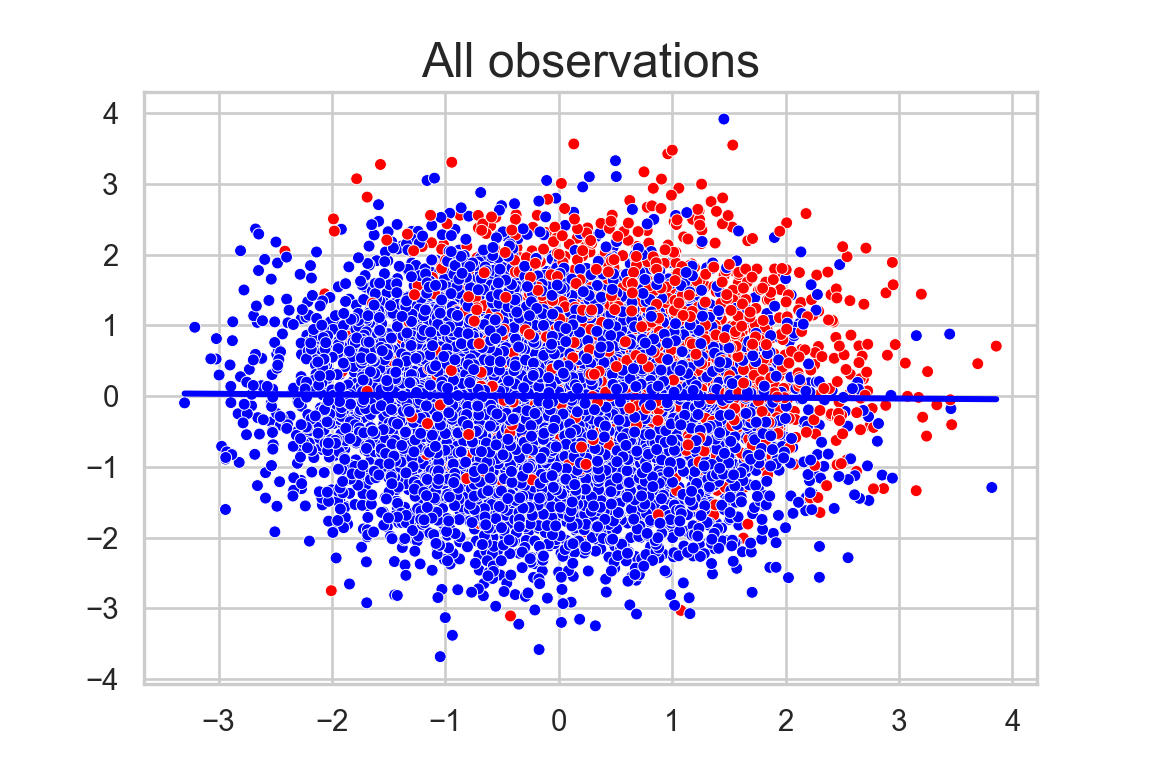
Number of observations (_N) was 0, now 10,000.Selection bias - All I
Call:
lm(formula = y ~ x, data = data_all)
Residuals:
Min 1Q Median 3Q Max
-3.9515 -0.6716 0.0087 0.6698 3.9878
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.011825 0.009994 -1.183 0.237
x -0.003681 0.010048 -0.366 0.714
Residual standard error: 0.9994 on 9998 degrees of freedom
Multiple R-squared: 1.342e-05, Adjusted R-squared: -8.66e-05
F-statistic: 0.1342 on 1 and 9998 DF, p-value: 0.7141Python
import statsmodels.api as sm
import pandas as pd
n = 10000
np.random.seed(100)
x = np.random.normal(size=n)
y = np.random.normal(size=n)
data1 = 1 / (1 + np.exp(2 - x - y))
group = np.random.binomial(1, data1, n)
data_we_see = pd.DataFrame({'x': x[group == 1], 'y': y[group == 1], 'group': group[group == 1]})
data_all = pd.DataFrame({'x': x, 'y': y, 'group': group})
X = data_all['x']
X = sm.add_constant(X)
y = data_all['y']
model = sm.OLS(y, X).fit()
print(model.summary()) OLS Regression Results
==============================================================================
Dep. Variable: y R-squared: 0.000
Model: OLS Adj. R-squared: 0.000
Method: Least Squares F-statistic: 1.281
Date: sex, 29 ago 2025 Prob (F-statistic): 0.258
Time: 13:31:33 Log-Likelihood: -14157.
No. Observations: 10000 AIC: 2.832e+04
Df Residuals: 9998 BIC: 2.833e+04
Df Model: 1
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
const -0.0003 0.010 -0.034 0.973 -0.020 0.019
x -0.0112 0.010 -1.132 0.258 -0.031 0.008
==============================================================================
Omnibus: 0.267 Durbin-Watson: 2.009
Prob(Omnibus): 0.875 Jarque-Bera (JB): 0.242
Skew: 0.009 Prob(JB): 0.886
Kurtosis: 3.017 Cond. No. 1.01
==============================================================================
Notes:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.Stata
Number of observations (_N) was 0, now 10,000.
Source | SS df MS Number of obs = 10,000
-------------+---------------------------------- F(1, 9998) = 0.28
Model | .284496142 1 .284496142 Prob > F = 0.5938
Residual | 9999.04347 9,998 1.00010437 R-squared = 0.0000
-------------+---------------------------------- Adj R-squared = -0.0001
Total | 9999.32797 9,999 1.0000328 Root MSE = 1.0001
------------------------------------------------------------------------------
y | Coefficient Std. err. t P>|t| [95% conf. interval]
-------------+----------------------------------------------------------------
x | -.0053101 .009956 -0.53 0.594 -.0248259 .0142057
_cons | .0006182 .0100006 0.06 0.951 -.0189849 .0202213
------------------------------------------------------------------------------Selection bias
Selection bias não é o único dos nossos problemas, mas é um importante.
Veja que suas conclusões mudaram significativamente.
Não seria difícil criar um exemplo em que o coeficiente verdadeiro fosse positivo.
Exemplo desses problemas

Source: Angrist
Não podemos pegar dois caminhos.
Exemplo desses problemas
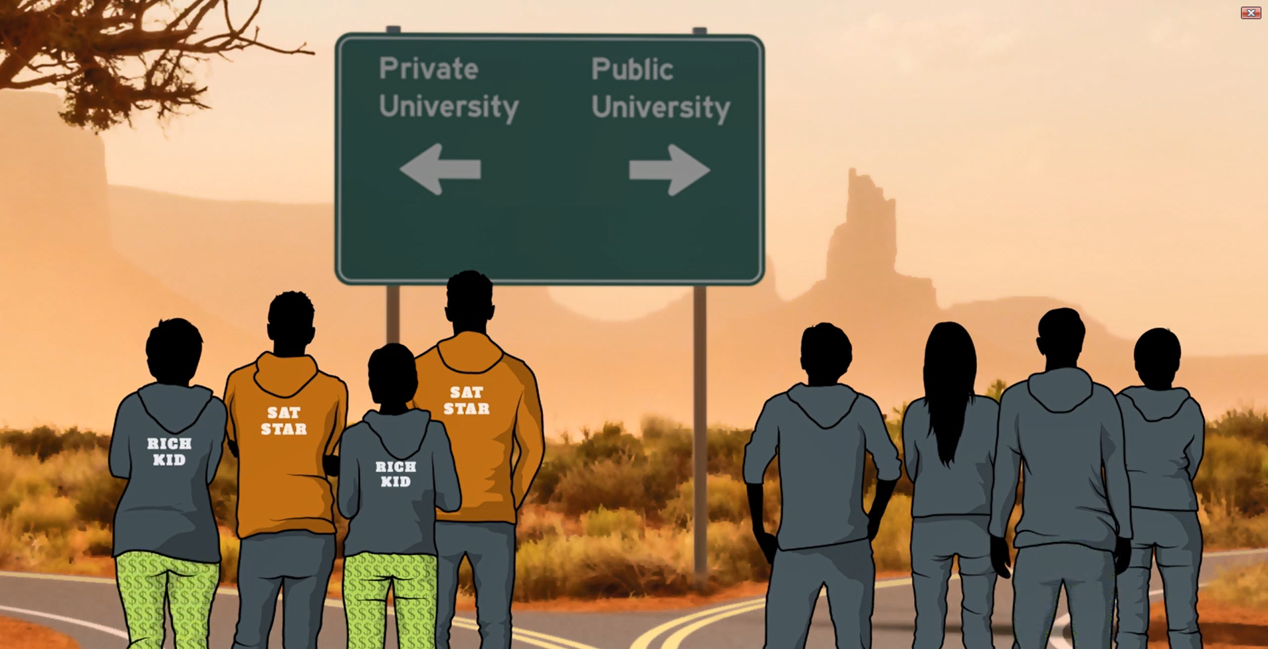

Source: Angrist
Não podemos comparar pessoas que não são comparáveis.
O que precisamos fazer?
Definir um bom Design empírico
No mundo ideal: teríamos universos paralelos. Teríamos dois clones, em que cada um escolhe um caminho. Todo o resto é igual.
- Obviamente, isso não existe.
Segunda melhor solução: experimentos
Mas o que é um experimento?
Grupo de tratamento vs. Grupo de controle
Igualdade entre os grupos (i.e., aleatoriedade no sampling)
- Nada diferencia os grupos a não ser o fato de que um indivíduo recebe tratamento e o outro não
- Estamos comparando maças com maças e laranjas com laranjas
Testes placebo/falsificação.
🙋♂️ Any Questions?
Thank You!


Henrique C. Martins [henrique.martins@fgv.br][Do not use without permission]